Determining an optimum determination mannequin is a vital however troublesome combinatorial process in imbalanced microarray-based most cancers classification.
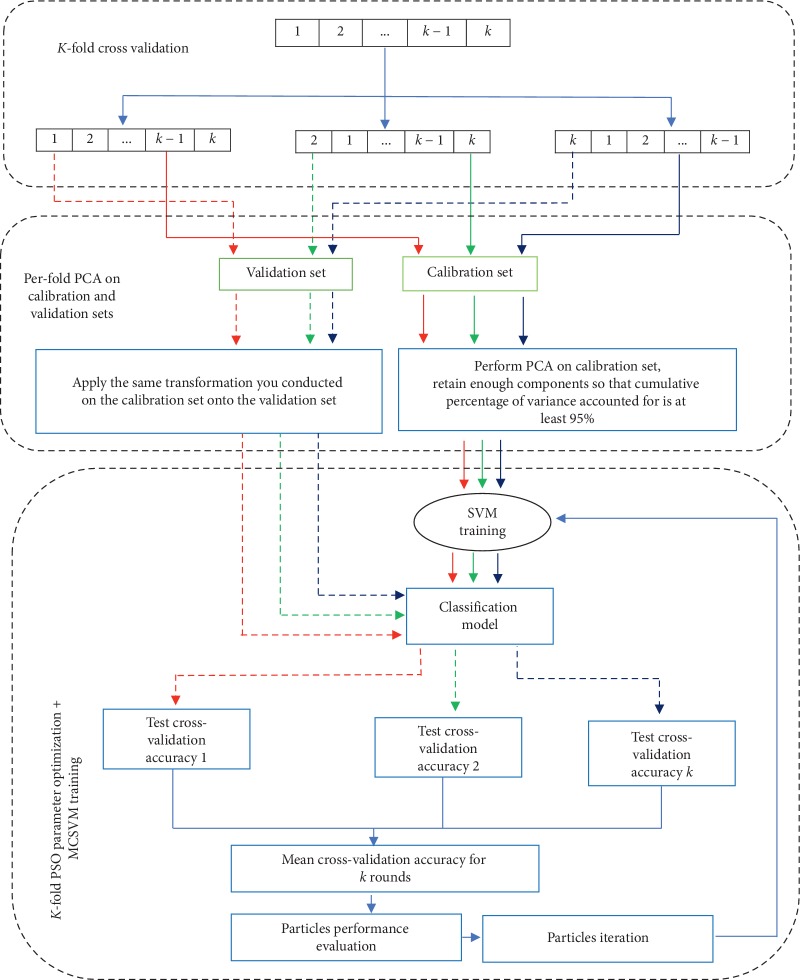
Though the multiclass help vector machine (MCSVM) has already made an essential contribution on this subject, its efficiency solely is determined by three elements: the penalty issue C, the kind of kernel, and its parameters. To enhance the efficiency of this classifier in microarray-based most cancers evaluation, this paper proposes PSO-PCA-LGP-MCSVM mannequin that’s based mostly on particle swarm optimization (PSO), principal part evaluation (PCA), and multiclass help vector machine (MCSVM).
The MCSVM relies on a hybrid kernel, i.e., linear-Gaussian-polynomial (LGP) that mixes some great benefits of three customary kernels (linear, Gaussian, and polynomial) in a novel method, the place the linear kernel is linearly mixed with the Gaussian kernel embedding the polynomial kernel. Further, this paper proves and makes certain that the LGP kernel confirms the options of a legitimate kernel.
In order to disclose the effectiveness of our mannequin, a number of experiments had been performed and the obtained outcomes in contrast between our mannequin and different three single kernel-based fashions, particularly, PSO-PCA-L-MCSVM (using a linear kernel), PSO-PCA-G-MCSVM (using a Gaussian kernel), and PSO-PCA-P-MCSVM (using a polynomial kernel).
In comparability, two twin and two multiclass imbalanced customary microarray datasets had been used. Experimental outcomes when it comes to three prolonged evaluation metrics (F-score, G-mean, and Accuracy) reveal the superior world characteristic extraction, prediction, and studying talents of this mannequin towards three single kernel-based fashions.

Identification of breast cancer-related circRNAs by evaluation of microarray and RNA-sequencing information: An observational examine.
An growing variety of research point out that round RNAs (circRNAs) take part in tumorigenesis. The goal of this examine was to elucidate the regulatory mechanisms of circRNAs in breast most cancers based mostly on the development of the circRNA-related ceRNA community.The expression profiles of circRNAs, miRNAs, and mRNAs had been obtained from the Gene Expression Omnibus (GEO) and The Cancer Genome Atlas (TCGA) databases.
A ceRNA community was constructed by Cytoscape. The interactions amongst proteins had been analyzed utilizing the STRING database, and hub genes had been extracted utilizing the cytoHubba utility.
The capabilities of the differentially expressed mRNAs (DEmRNAs) had been analyzed by the Kyoto Encyclopedia of Gene and Genomes (KEGG) and the Gene Ontology (GO) database.In complete, 7 differentially expressed circRNAs (DEcircRNAs), 27 differentially expressed miRNAs (DEmiRNAs), and 102 DEmRNAs had been chosen for the development of the ceRNA community of breast most cancers. We established a protein-protein interplay community and recognized 6 hub genes.
Then, a circRNA-miRNA-hub gene regulatory module was established based mostly on 2 DEcircRNAs, 2 DEmiRNAs, and a pair of DEmRNAs. GO and KEGG pathway analyses indicated the attainable affiliation of DEmRNAs with breast most cancers onset and development.The circRNA hsa_circ_0000519 is probably going important within the pathogenesis of breast most cancers and will function a future therapeutic biomarker.